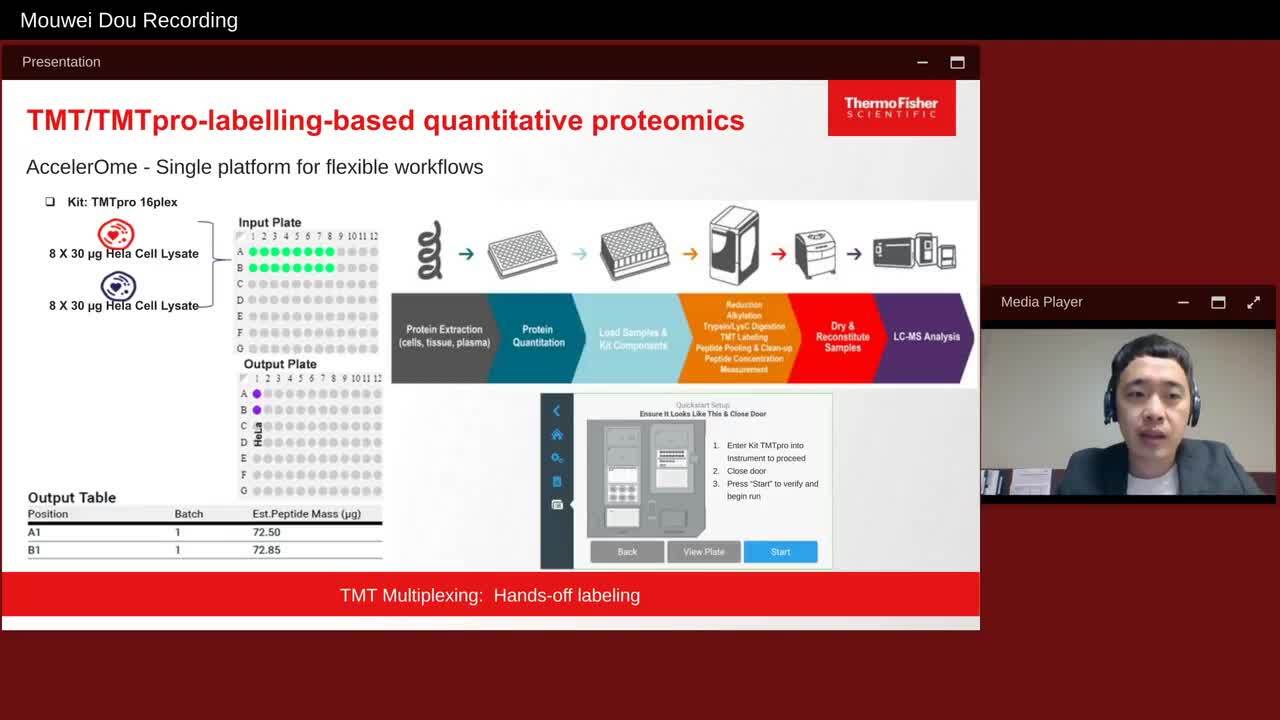
TMT: A Versatile and Powerful Approach for Characterization of Proteomes
40:02
Isobaric labeling TMT experiments with optimal depth and quantitative accuracy for Proteomics.
Related Videos
In Mass Spectrometry Videos
-
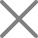
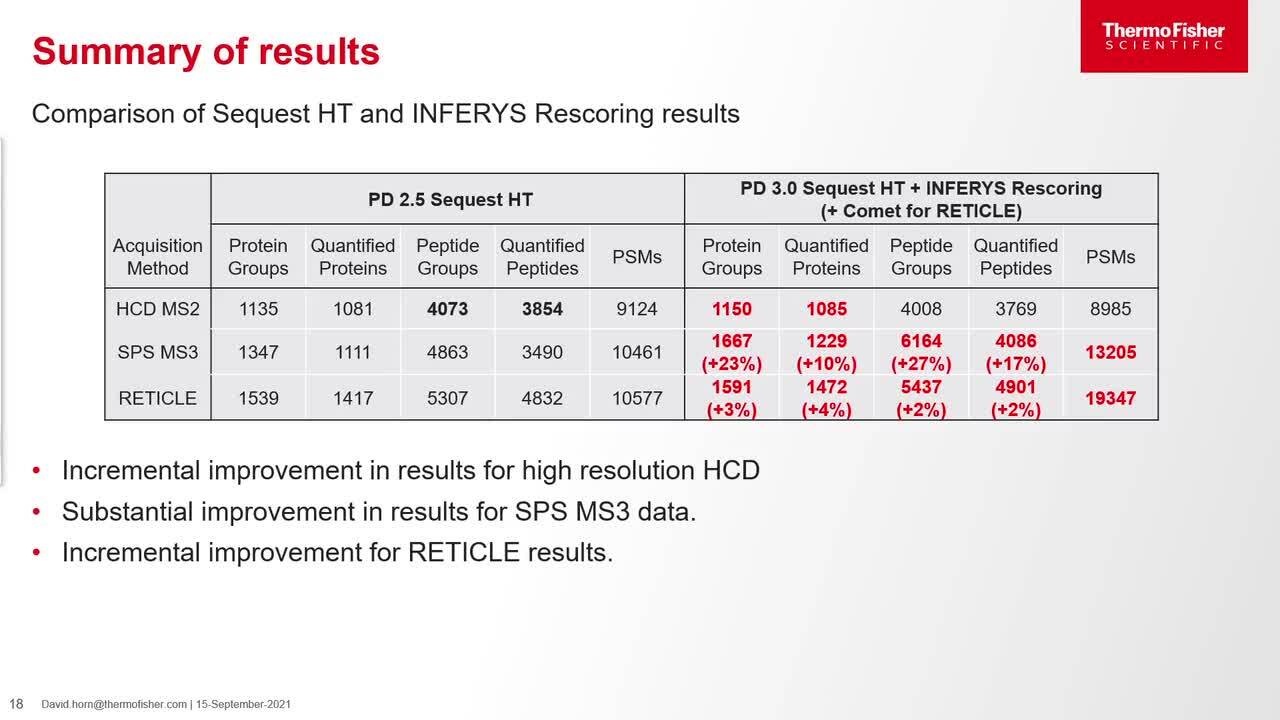
Play video Deep learning tools in PD 3.0 (CHIMERYS and INFERYS) for TMT analysis
Deep learning tools in PD 3.0 (CHIMERYS and INFERYS) for TMT analysis
Deep learning tools in PD 3.0 (CHIMERYS and INFERYS) for TMT analysis
31:42
-
Play video Measuring Protein-Turnover with Heavy Isotope Labeling and Complement Reporter Ions
Measuring Protein-Turnover with Heavy Isotope Labeling and Complement Reporter Ions
Measuring Protein-Turnover with Heavy Isotope Labeling and Complement Reporter Ions
29:34
-
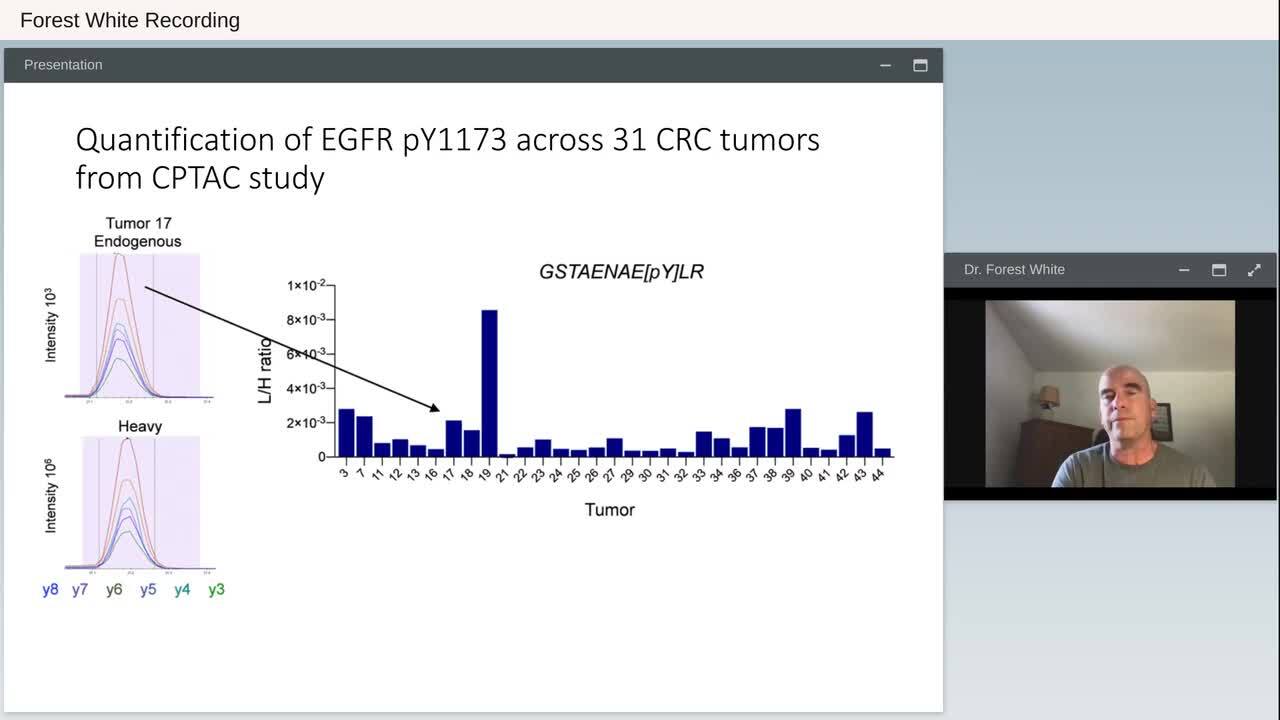
Play video Clinical Phosphoproteomics to Uncover New Therapeutic Strategies
Clinical Phosphoproteomics to Uncover New Therapeutic Strategies
Clinical Phosphoproteomics to Uncover New Therapeutic Strategies
26:22
-
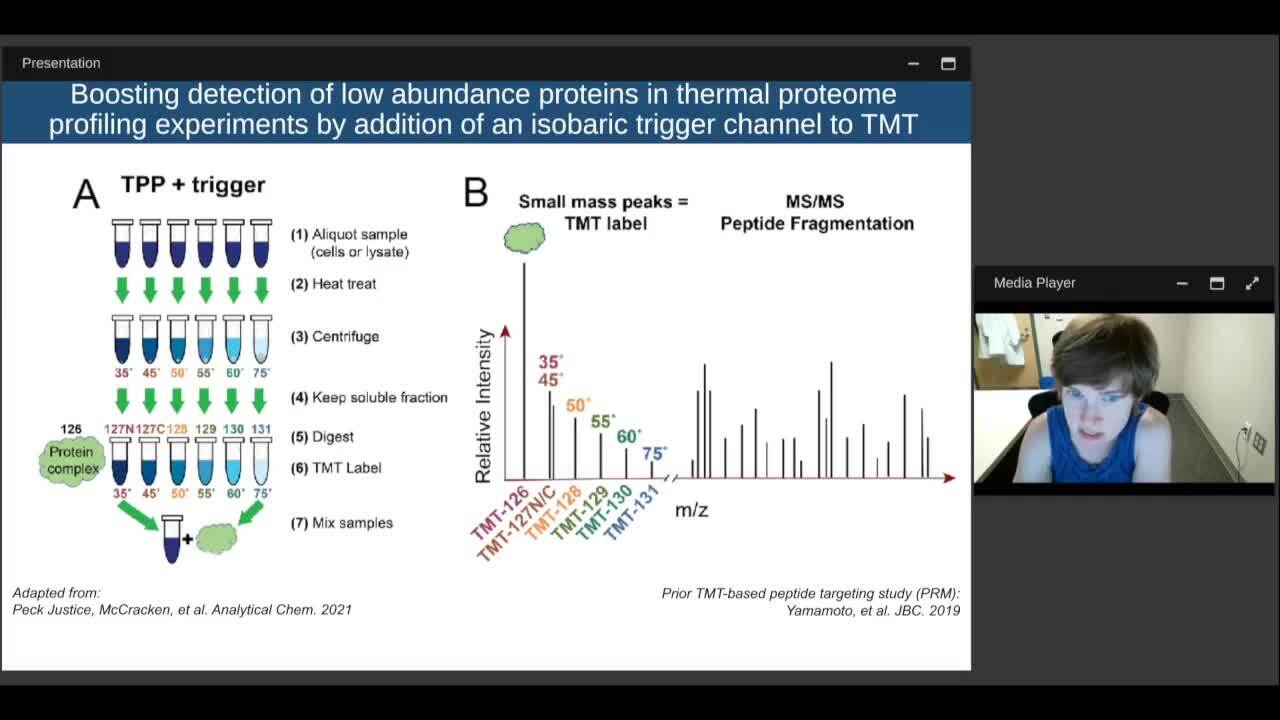
Play video Thermal Proteome Profiling for Functional and Molecular Characterization of Variant Proteins
Thermal Proteome Profiling for Functional and Molecular Characterization of Variant Proteins
We propose the use of thermal proteome profiling (TPP) for studying the impact of these disease related sequence variants on a systems wide level. TPP is a bottom up, TMT-based, quantitative mass spectrometry technique for determining the melt curves
30:52
-
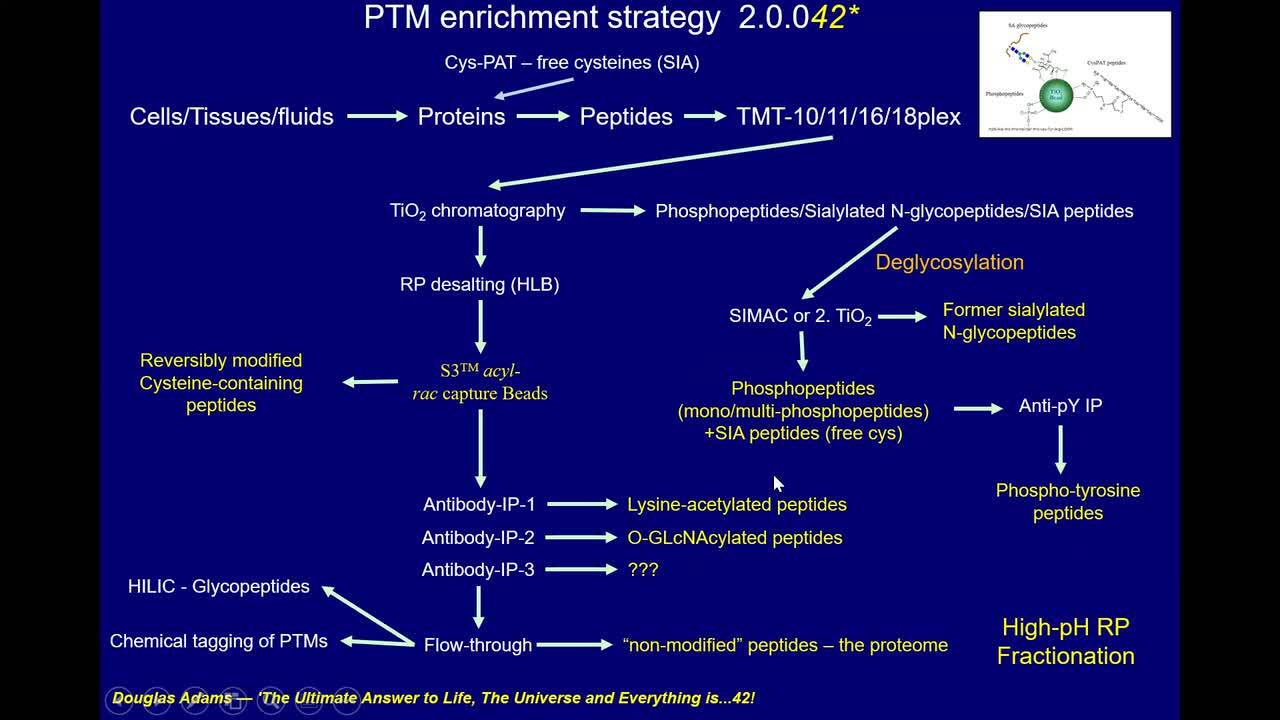
Play video Modulation of Cysteines upon cellular signalling
Modulation of Cysteines upon cellular signalling
In the present talk we will introduce cysteine modifications, various tools for assessing these and various applications were we have used cysteine-specific proteomics to characterize important cysteine residues in key proteins for cellular signallin
37:56
-
Play video An Automated Sample Preparation Solution for Mass Spectrometry-based Proteomics
An Automated Sample Preparation Solution for Mass Spectrometry-based Proteomics
The automated platform enables hands-off protein lysis, DNA removal, protein reduction and alkylation, digestion, TMT labeling, pooling and cleanup, and peptide concentration measurement, providing several workflows and companion reagents to perform
18:25